Know in one minute about EcoRI Restriction Enzyme
|
Introduction
EcoRI is defined as a restriction enzyme that acts as an essential tool in molecular biology research. Restriction enzymes belong to a class of enzymes called nucleases. They are molecular scissors for manipulating DNA. Hence are responsible for cutting DNA at specific sites known as recognition sites.
- Endonucleases that remove nucleotides from the ends of DNA.
- Exonucleases that cut the DNA at specific positions anywhere within its length.
EcoRI is isolated from the prokaryotic cell of Escherichia coli and is an example of a restriction endonuclease enzyme that cuts DNA at specific sites.
History of EcoRI
- EcoRI is a type II restriction enzyme that was isolated in H.Boyer’s lab by Stewart Linn and Werner Arber in 1963.
- They are responsible for restricting the growth of bacteriophages.
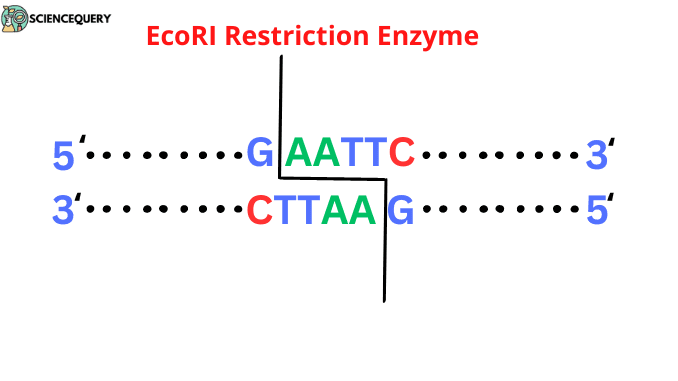
- It generally cleaves at a 5′-GAATTC-3′ sequence.
- The recognition site of EcoRI is a palindrome sequencing site. Palindromes are a group of letters that form the same when reading both forward and backward i.e., 5′-3′ or 3′-5′ direction.
Structure of EcoRI
- EcoRI has a homodimeric structure and a molecular mass of 31kDa.
- The structure of EcoRI is mostly palindromic along with 4-8 base pairs.
- Each type II restriction enzyme has its specific restriction site and cleavage properties.
- As per the structure is concerned, EcoRI has a dimeric resolution.
- Every subunit consists of a catalytic active domain and a recognition domain that is an active site of binding.
Enzyme Composition
EcoRI is extensively used in genetic engineering and contains about 377 amino acids. These acids play an important role in cleaving DNA. The binding site of the enzymes binds only by DNA ligase.
- The cleavage of DNA takes place in the presence of Mg²+ ions or near recognition sites.
- It enables sticky ends that are produced at the ends of DNA and facilitates the action of the enzyme DNA ligase.
- The sticky ends are ligated to the complementary DNA strand.
The active site of EcoRI contains four amino acids namely histidine (H), aspartic acid (D), arginine (R), and tyrosine (Y). The Mg²+ ion helps keep this structure stable by surrounding the phosphate backbone and balancing the interactions between elements such as hydrogen bonds at the base pair edges.
Active Site of EcoRi Enzyme
- The active site of Ecori contains amino acid residues such as Asp767, His386, and Ser338 that act as sites for catalysis.
- At this active site, these amino acid residues bind with the DNA molecules, thus matching the recognition sequence.
- When two complementary strands are joined, they form a duplex due to the presence of ‘H-bonds’ between them.
Mechanism of action
-
Specificity of EcoRI
1. It has high specificity as it binds to only certain sequences of nucleotides called recognition sites.
2. This specificity makes it useful in genetic engineering and biotechnology research.
3. It allows researchers to keep track of gene alterations, hence facilitating discoveries.
-
Cutting and Cleaving
1. Cutting and Cleaving DNA molecules is an essential technique to break long strands of DNA into small fragments.
2. A commonly used enzyme is EcoRI Restriction endonucleases, which act as molecular scissors on dsDNA.
Hence, it allows scientists a better understanding of the working mechanism of a gene.
-
Formation of Sticky Ends
1. The formation of sticky ends by EcoRI is a process used in DNA cloning.
2. When DNA sequences are cut by restriction endonucleases, the two ends formed are called ‘Sticky ends’.
Applications of EcoRI
1. DNA fragmentation & Cloning
EcoRI plays a key role in the world of molecular biology. The fragmentation is done by keeping incubated the purified DNA at optimal pH and temperature.
Cloned genes are used as probes and help in disease diagnosis.
2. rDNA technique
This technique is used for the production of safe and therapeutic drugs that prove to be a savior to mankind.
ADA deficiency has proven to be cured if a functional gene is introduced at an early embryonic stage in bone marrow cells.
EcoRI in Genetic Mapping
The determination of a base sequence in a fragment is called DNA sequencing. Southern hybridization involves the separation of DNA by gel electrophoresis. The DNA is denatured before blotting.
EcoRI in Genomic Studies
Gene mapping is a schematic representation of various genetic markers. It is arranged in an orderly fashion. They are also known as linkage maps as they measure the recombination frequencies and differences between markers.
RFLP and SNP (single nucleotide polymorphism) are widely used in genomic studies.
Variants of EcoRI
EcoRI is a DNA-cutting enzyme and has several variants. It includes MLU I, EcoRI-BamHI, EcoRI-Xhoi, and GaBI SI WT PhoA HIS6 Restriction endonucleases. These variants are useful in determining specificity in genome sequencing projects.
Advantages of EcoRI
- EcoRI cuts DNA at the exact location of interest.
- Specificity is high.
- It eases out gene cloning and genomic sequencing applications.
- It promises stability and the ability to withstand higher temperatures, allowing smooth research.
Disadvantages of EcoRi
- Advancement in restriction enzymes is necessary to get verified results.
- Cost efficiency is high.
- Result analysis is time-consuming.
Q&A
What is the function of EcoRi?
EcoRI is a restriction endonuclease, extensively used in molecular biology projects to cleave DNA at specific positions and generation of sticky ends for easy ligation.
Why is it called EcoRI?
It is an example of type II restriction endonuclease extracted from E. coli bacteria. ‘R’ denotes the specific strain- RY13 and I denote the 1st enzyme to be derived from this particular strain.
How does EcoRI cut DNA?
This restriction enzyme cuts DNA at the GAATTC site, where G stands for guanine, A stands for Adenine, T stands for thymine, and C for cytosine.
What is EcoRI cleaving of DNA?
The cleaving site of DNA starts from 5′ GAATTC 3′ sites that consist of 6 base pairs on a nucleotide sequence.
References
- https://academic.oup.com/nar/article-abstract/42/1/3/2438402
- https://pubs.acs.org/doi/pdf/10.1021/bi00002a037
- https://www.future-science.com/doi/abs/10.2144/btn-2021-0083
- https://journals.asm.org/doi/abs/10.1128/mcb.14.7.4493-4500.1994
- https://www.nature.com/articles/s41598-020-73321-8
Written By: Sushmita Mukhopadhyay
